AI-Driven Linear Peptide Optimization and Design Platform
a. Product Overview
This product is an integrated, cutting-edge platform for linear peptide optimization and de novo design, powered by artificial intelligence and computational structural biology. The platform, AlanPepAI™, accepts an initial peptide sequence (or works solely from a target protein structure) and, through fully automated AI iterative optimization and physical modeling, delivers a series of novel linear peptide candidate molecules within days. These candidates exhibit significantly enhanced binding affinity and optimized physicochemical properties, greatly accelerating the progression from hit to lead compound.
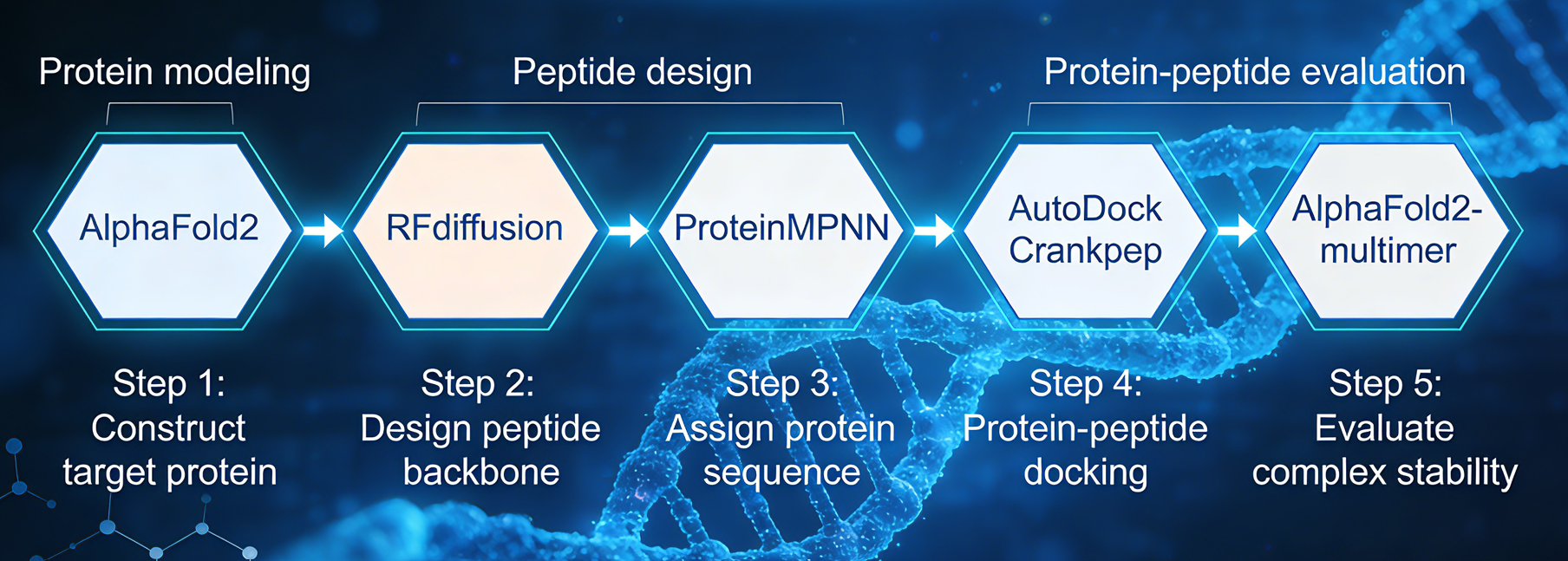
The core technology integrates next-generation protein structure prediction (AlphaFold2), AI-driven intelligent sequence design (ProteinMPNN), and high-precision peptide–protein docking (AutoDock CrankPep) into a fully closed-loop design-validation workflow. Compared to traditional site-directed mutagenesis or limited library screening, this platform offers three key advantages: Innovation, as the AI explores sequence spaces beyond human experience to generate unique structural motifs with novel mechanisms of action and strong intellectual property potential; Systematic Capability, enabling multi-objective parallel optimization of individual or multiple performance indicators (e.g., affinity, stability, solubility); and Predictability, providing comprehensive virtual validation from atomic-level 3D complex structures to binding free energy, significantly increasing experimental success rates while reducing development costs and risks.
b. Core Advantages of AlanPepAI™
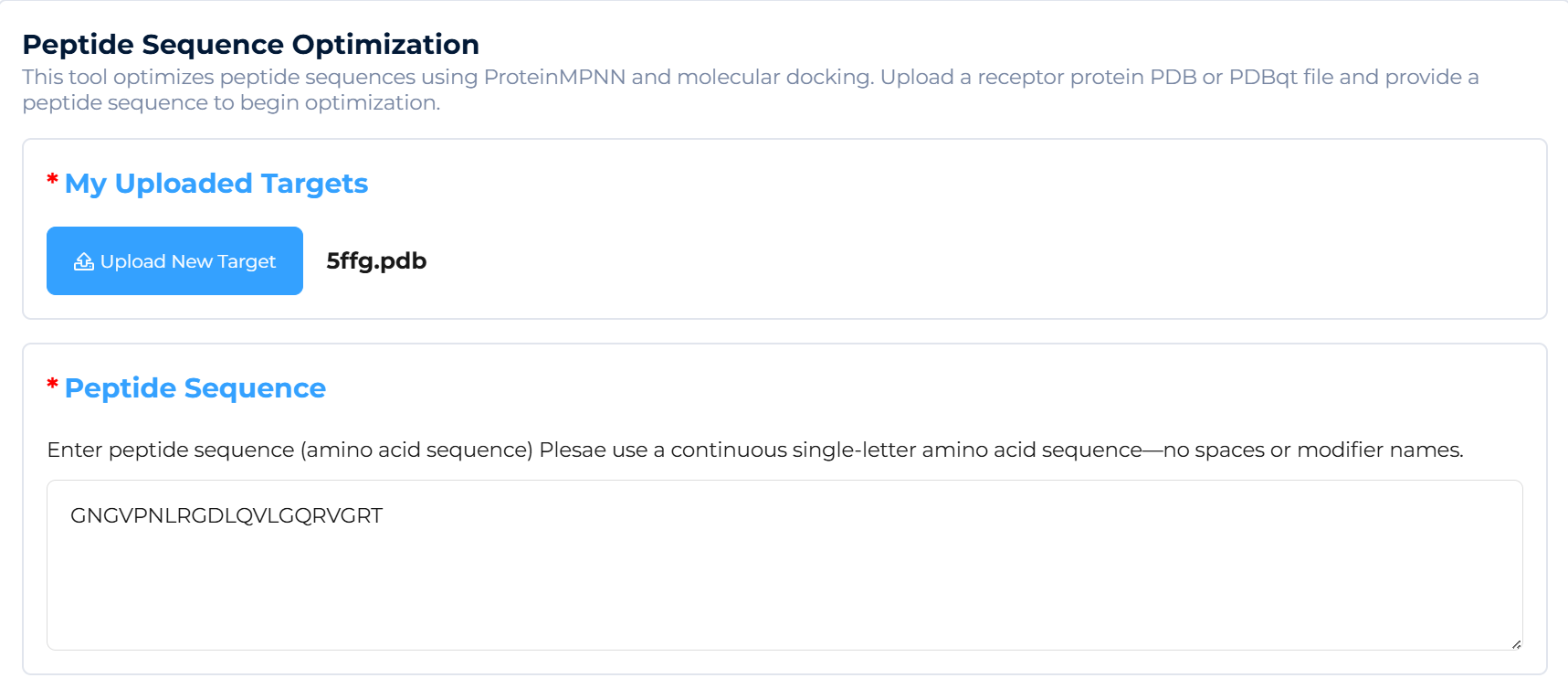
Requirement Analysis & Input Preparation: The client provides the three-dimensional structure file of the target protein (in PDB format) and the initial linear peptide sequence to be optimized (in single-letter amino acid code format). The platform automatically processes the structure file, identifies key binding sites, and analyzes the binding mode of the initial peptide, establishing a baseline for optimization.
Structure-Guided Sequence Optimization & Generation: The platform's core workflow initiates. First, it utilizes AlphaFold2 to accurately build or refine the structural models of the target protein and the initial peptide. Subsequently, ProteinMPNN, an advanced neural network for protein sequence design, performs large-scale, intelligent redesign of the initial peptide's amino acid sequence. This step is not random mutation; it is an inverse design process based on the 3D structure of the target-peptide complex, generating multiple novel candidate sequences that are structurally more complementary and have optimized interactions.
High-Precision Affinity Assessment & Screening: For the thousands of generated candidate sequences, the platform employs specialized tools like AutoDock Crankpep for rapid, precise molecular docking simulations. This predicts their binding conformation with the target and calculates binding affinity (scoring). Simultaneously, AlphaFold-Multimer can be integrated to validate the structural stability of high-scoring complexes. All candidate peptides are comprehensively ranked and filtered based on multi-dimensional metrics, including predicted binding scores and structural quality.
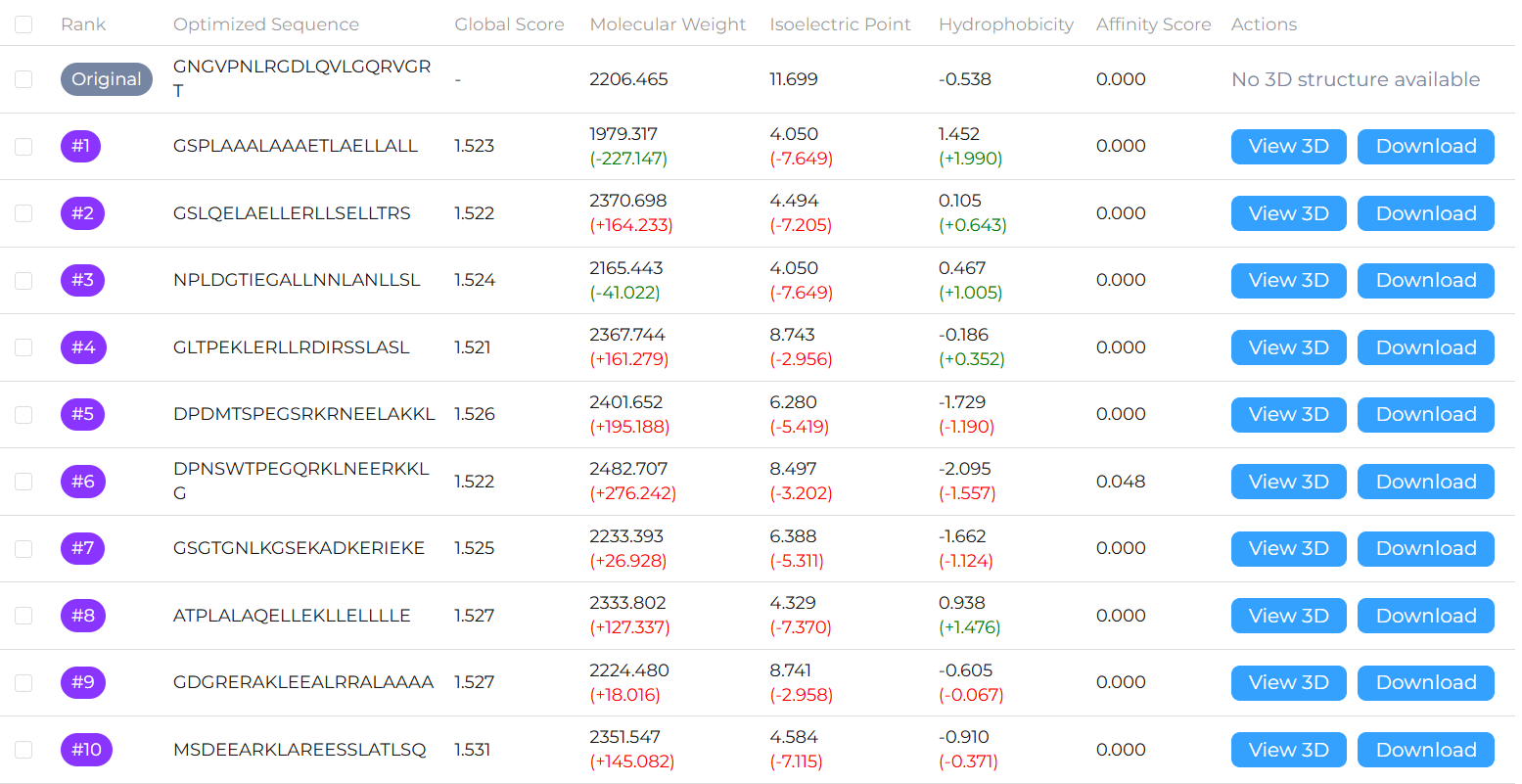
In-Depth Analysis & Results Delivery: The platform delivers in-depth interaction analysis for top-ranked candidate peptides (typically the Top 10), supported by high-resolution 3D binding structures for direct reference. It visually highlights critical target interactions, including hydrogen bonds and hydrophobic contacts, and generates a comprehensive property prediction report that supports confident, data-driven decision-making in downstream development.
c. For Whom and What Problems Does It Solve?
Challenges
Traditional peptide discovery relies on natural peptide libraries, synthetic peptide libraries, or phage display, which suffer from limited diversity, long development timelines, high costs, and poor in vivo stability of linear peptides.
Peptide sequences are highly hydrophobic, leading to high synthetic difficulty.
Poor peptide affinity towards targets.
Weak binding ability between the spatial conformation of polypeptides and their corresponding targets.
Solutions
The platform can generate a series of high-affinity candidate molecules with intellectual property novelty for specific targets (PDB or PDBqt file required) within minutes to hours, shortening the early discovery cycle from months to days.
d. Typical Workflow
Input: Submit the target protein sequence structure (PDB or PDBqt file).
Design: The diffusion-based workflow rapidly generates a diverse pool of high-affinity linear peptide sequences while simultaneously predicting their binding modes, enabling efficient identification and prioritization of the most promising candidates for experimental validation.
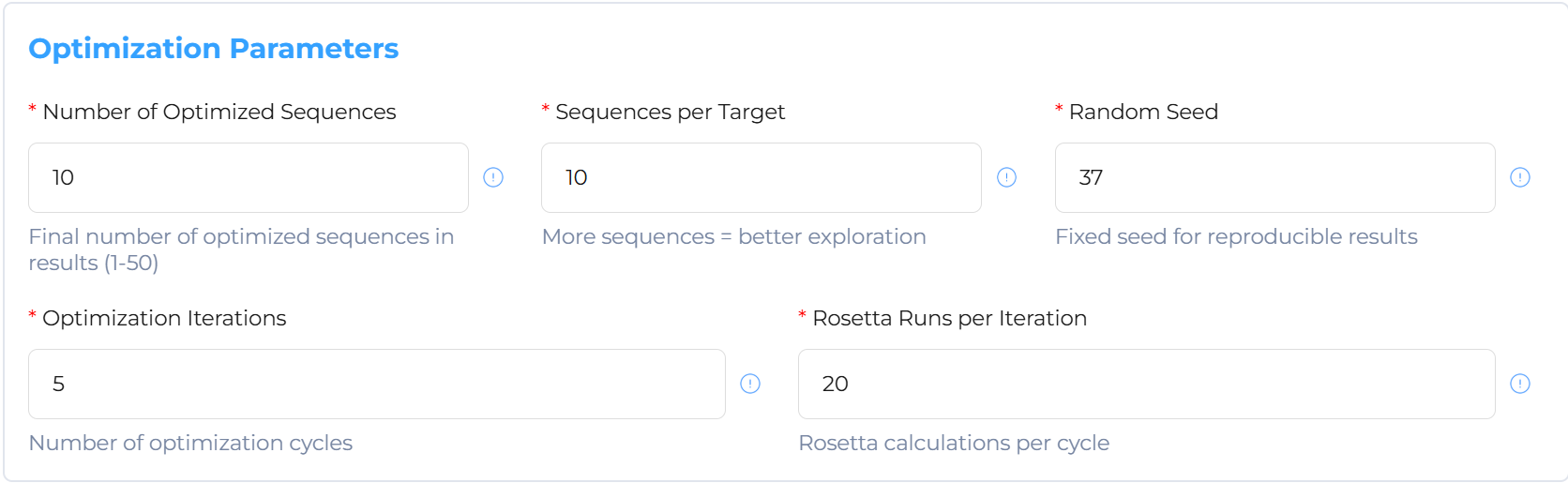
Optimization: Perform structure-based and AI-driven refined adjustments on top-tier sequences.
Output: Obtain a series of virtually validated linear peptide candidates and their predicted complex structures (pdbqt files with 3D viewer platform) for subsequent synthesis and testing.
e. Cost
Each optimization project costs 100 credits
f. Showcase
![]()
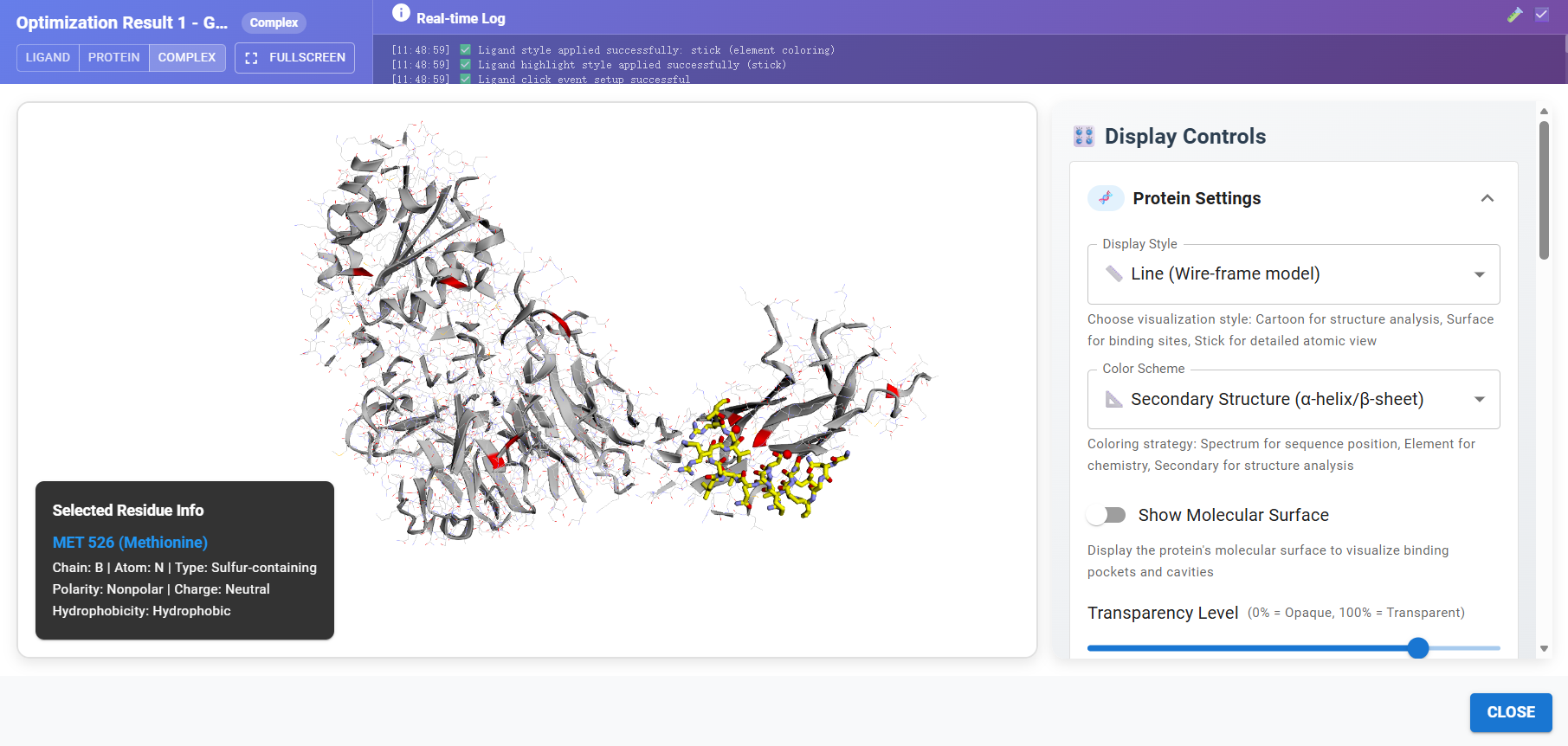
Other AI-Powered Quantum Computing Tool Platforms:
AlanMolecularAI™: AI-Quantum Computing-Powered Online Optimization Platform for Small Molecules, integrated with proprietary molecular fragmentation technology, enables multi-round ADMET model screening and accelerates the early discovery and optimization of small molecules
AlanDockAI™: Online Tool for Automatic Small Molecule-Target Docking with Proprietary Algorithms
To Get More Credits or View Pricing Plans:
Log in and go to My Account > AIPower Platform > Add Credits to Your Account
For Peptide Synthesis Service:
Please click here Peptdie Synthesis